Time-Varying Temporal Functional Modes: A Novel Model to Probe Time-Varying Functional Reconfigurations in fMRI While Accounting for Shared Anatomical Infrastructure
Tamara de Kloe1,2
Supervisors: Alberto Llera Arenas1,2, Nils Kohn1,2, Christian F Beckmann1,2,3
1: Donders Institute for Brain, Cognition and Behaviour
Radboud University, Nijmegen, The Netherlands
2: Department of Cognitive Neuroscience
Radboud University Medical Centre, Nijmegen, The Netherlands
3: Wellcome Centre for Integrative Neuroimaging
FMRIB Building, Nuffield Department of Clinical Neurosciences
University of Oxford, Oxford, United Kingdom
Author Note
Corresponding author: de Kloe, Tamara Jedidja, t.dekloe@donders.ru.nl
Abstract [back to top]
The core cognitive neuroscientific aim is to determine how the brain gives rise to mental function. The answers provided to this ontological question depend in part on predominant methodology. Initial functional magnetic resonance imaging (fMRI) studies were mainly segregationist, that is, focused on mapping psychological functions to individual brain sites. The modularity hypothesis underlying this approach, however, has been questioned since its conception. Conversely, it is argued that mental function relies on interactions within and between integrated functional brain networks. This is, for example, studied using functional connectivity methods, which examine co-variation of spatially distributed signals emitted from the brain. Time-averaged estimates of functional connectivity are commonly obtained. A recent line of research, however, aims to probe time-varying changes in connectivity, based on the premise that biologically meaningful changes in inter-regional relations are expected to occur. In this work, we present a novel time-varying functional connectivity method that explicitly accounts for spatial overlap, that is, for functional networks to share a common anatomical infrastructure. The model is an extension of the work by Smith et al. (2012), in which a spatial decomposition and a temporal independent component analysis are applied consecutively, resulting in functionally distinct and spatially overlapping networks called "Temporal Functional Modes" (TFMs). The extent to which TFMs recruit spatial parcels (described by the TFM mixing matrix) remains time-invariant in the original model. Here, we introduce a time-resolved version of TFMs to account for the time-varying nature of functional connectivity. We show analytically that the fixed mixing matrix in classical TFM analysis is the temporal average of a time-varying mixing matrix which can be obtained in closed form and which represents moment-by-moment estimations of brain network reconfigurations. We apply the novel method to a high-temporal resolution fMRI dataset and demonstrate that the retrieved network reconfigurations follow the expectation based on the task design.
Keywords: time-varying functional connectivity, dynamic functional connectivity, independent component analysis, temporal functional modes, spatial overlap
Introduction [back to top]
Much like glasses through which one views the world, the choice of method critically determines the questions that can be asked and the limits of what can be observed in studying brain function. In the last few decades, neuroimaging has become an essential tool for studying the temporal dynamics and spatial topology of the brain in vivo. The former is usually studied using electroencephalography (EEG), and for the latter, functional magnetic resonance imaging (fMRI) is most common due to their relatively high temporal and spatial resolution, respectively (Bandettini, 2009; Rosa et al., 2010). Initial fMRI studies probed the neural substrates of brain function within a segregationist paradigm, focusing on mapping psychological function to individual activation sites. The corresponding methodological approach consists of per-voxel comparisons of blood oxygen level-dependent (BOLD) signals to the expected time series, based on experimental designs constructed to single out a cognitive function. Statistical tests on these comparisons provide maps that show what parts of the brain are associated with that particular cognitive function (Poldrack et al., 2011; Smith et al., 2004). Inherent to this mass-univariate approach is that voxels are studied independently, hence precluding examination of integration or collaboration. Since the discovery that spatially distinct and temporally synchronized brain regions can be differentiated in fMRI (Biswal et al., 1995), the field has started to characterise these integrated functional networks. A complementary integrationist approach is adopted, focused on characterizing integrated, spatially distributed functional brain networks and their role in cognition (Eickhoff & Muller 2015). Methods for Studying Integration in fMRI Integrated functional brain networks can be probed using functional connectivity methods. Popular methods such as seed-based correlation analysis (SCA; Biswal et al., 1995) and spatial independent component analysis (ICA; Beckmann and Smith, 2004; Beckmann, 2012; McKeown and Sejnowski, 1998) have reliably differentiated these networks termed resting-state networks across participants (Beckmann et al., 2005; Damoiseaux et al., 2006), both in rest and during tasks (Smith et al., 2009). Alterations in functional connectivity of large-scale brain networks have proven to relate to cognition (e.g., Laird et al., 2011), behaviour (Rosazza and Minati, 2011), human development (Uddin, Supekar & Menon, 2010), and personality (Mulders et al., 2018). Moreover, they show promise as a biomarker, as they relate to a variety of pathologies such as depression (Mulders et al., 2015; Greicius et al., 2007), schizophrenia (Lynall et al., 2010), and Alzheimer’s disease (Damoiseaux et al., 2012; Greicius et al., 2004; Lehmann et al., 2015; Lustig et al., 2003). Functional connectivity is conceptually described as communication between brain areas to support cognitive function. Methodologically, it concerns statistical association between spatially distinct brain regions (Friston et al., 1993). The first described method, SCA, obtains correlations between a previously defined region of interest and all other parts of the brain. This allows for hypothesis-driven analyses, for example, probing whole-brain connectivity of an area that has previously been related to a particular cognitive function. This approach, however, restricts the analysis to one chosen region or system, although connectivity changes between other regions might occur (Bijsterbosch, 2017). The second method, spatial ICA, is a multivariate and model-free blind source separation technique that assumes that the fMRI signal consists of a linear combination or mixture of signals from different sources. The signals are decomposed ("unmixed") into a set of spatial components that are statistically maximally independent by extracting non-Gaussian sources (Beckmann and Smith, 2004; Beckmann et al., 2005; Beckmann, 2012). The extracted components are generally understood as brain networks that show similar BOLD activity (McKeown, 2003). The data can also be decomposed in temporally independent sources using temporal ICA. Of the two, spatial ICA is more common in fMRI. An important reason is that in fMRI, the spatial dimension is generally much larger than the temporal dimension (i.e., more voxels than volumes are obtained; Smith et al., 2012; Calhoun et al., 2001). In either case, the sources are described by means of a spatial map and a corresponding time series. Both these spatial and temporal features can subsequently be related to behaviour or be compared between certain groups of individuals. Almost all available ICA-based functional connectivity studies are performed in resting-state. In part, because this does not provide a behavioural benchmark to which the time series can be related like in task-based fMRI, the gold standard is to statistically compare the spatial maps across individuals or groups, for example, to inform on significant differences in strength, shape or location of the networks (Bijsterbosch et al., 2018). Time-varying functional connectivity
Rationale [back to top]
Most functional connectivity methods assume that the dependence structure between brain regions is constant over the course of an fMRI scan or experimental condition. In SCA, for example, average correlations between BOLD signals of specific seed and target regions over an entire scan duration or condition are used as a proxy for the network behaviour. In spatial ICA, the spatial map weights, which reflect the connection strengths between regions, are constant over time (Hutchison et al., 2013). Many significant advances have been made in understanding large-scale brain networks while working under this assumption. A recent line of research, however, posits that it fails to capture the expected changes in interactions between, or shifts in dominance of, large-scale brain networks (Liégeois et al., 2017, 2019; Lurie et al., 2020; Preti et al., 2017). It is argued that characterisation of time-varying functional connectivity metrics is required, that is, time-varying changes in integration of spatially distributed functional brain networks should be examined. The term often used in the literature is "dynamic functional connectivity" (Preti et al., 2017). In a recent joint article by various leading experts in this developing field (Lurie et al., 2020), it was argued that standardisation of terminology is paramount. "Time-varying functional connectivity" (TVFC) was proposed, as differences between disciplines in use and definitions of the term "dynamic" can cause ambiguity. Conceptually, TVFC is defined as functional connectivity that varies as a function of time, and hence concerns temporal reconfiguration of functional entities (Iraji et al., 2020). Methodologically, it means that the parameters used to describe the dependence structure between these functional entities vary across time. It was pointed out that TVFC is broader than the static-dynamic distinction, whereas this has previously (mistakenly) been taken to mean the same thing (Lurie et al., 2020). Moreover, it has been shown that certain TVFC processes fall within the space of stationary models (Liégeois et al., 2017). Thus, not every TVFC metric implies non-stationarity, which is defined by a non-constant mean and covariance (or other higher-order moments; Hutchison et al., 2013). Time-varying functional connectivity methods. A range of TVFC methods has been suggested in recent years, which differ in several aspects. First, a distinction can be made between methods that explicitly model the underlying neural processes (i.e., dynamic biophysical models) and those that characterise the time-resolved dependence structure from the BOLD signals. A first important differentiation between methods of the latter category is the temporal window considered, ranging from individual frames or time points (i.e., instantaneous connectivity) to larger time windows (Preti et al., 2017). The functional connectivity metrics that are obtained per time point or window are either directly summarised using graph-theoretic measures, or "brain-states" are estimated using matrix factorisation techniques (e.g., k-means clustering or principal component analysis). These brain states are subsequently described using time-varying parameters such as transition probabilities and dwell times (Lurie et al., 2020; Preti et al., 2017). Sliding-window approaches. Most methods that characterise the time-resolved dependence structure directly from the BOLD signals are variations of the sliding-window pairwise correlation approach (for a review see Preti et al., 2017). In its most rudimentary form, this consists of segmenting the entire scan duration into smaller windows and computing a functional connectivity metric over successive (overlapping or non-overlapping) windows (Preti et al., 2017). Many extensions of this framework have been proposed, aiming to deal with the strong objection that window type and length have a substantial impact on the characterisation of the functional dynamics. On the one hand, small windows can introduce spurious correlations because of insufficient information to estimate TVFC robustly (Leonardi & Van De Ville, 2015). Given the inverse relationship between periodicity and frequency, another problem is that short windows, by definition, only allow probing high frequencies. Although there is some evidence that resting-state networks are broadband processes (Niazy et al., 2011; van Oort et al., 2012), they are generally assumed to be dominated by low-frequency oscillations (around .015Hz; Niazy et al., 2011), which will appear constant for short time windows. On the other hand, large windows smooth out the time-varying properties of interest (Hindriks et al., 2016; Iraji et al., 2020). A multivariate, ICA-based approach to sliding-window analysis has also been suggested (Kiviniemi et al., 2011). Performing spatial ICA on a sliding-window basis results in a set of independent components per window. Using this approach to analyse Default Mode Network (DMN) activity, Kiviniemi et al. (2011) concluded that DMN subnetworks dynamically change interactions throughout a full acquisition length and that activity of the whole DMN (as identified by stationary ICA) never occurs. Splitting of the DMN into multiple sub-networks, however, depends on the dimensionality of the ICA. A second objection to this multivariate ICA-based sliding window approach is that components need to be matched across windows, which is problematic since the order of components extracted using ICA is relatively arbitrary (Bijsterbosch, 2017) Time-frequency approaches. The seminal approach in electrophysiology called time-frequency analysis has also been applied to TVFC. It explores connectivity at multiple frequencies and circumvents the described window constraints of sliding window approaches. The seminal time-frequency approach to TVFC in fMRI is Wavelet Transform Coherence (Chang & Glover, 2010), where the size of the "window", that is, the wavelet, is adjusted based on the time scale of the frequencies under inspection. Time-frequency analyses result in a description of the amount of joint power between two time series and their relative phase as a function of time and frequency. Using this approach, it has been found that within and between-network connectivity patterns differ across frequencies, suggesting the presence of overlapping interactions that are potentially averaged in classical approaches (Chang & Glover, 2010; Yaesoubi et al., 2015). A longstanding idea is that low frequency, slow-wave signals drive resting-state networks. Probing their spectral characteristics using a Hilbert-Huang transform, Niazy et al. (2011) found that although low-frequency signals indeed dominate resting-state networks, they are characterised by different frequencies at different levels of phase synchrony. That is, they were found to be broadband processes. Leading Eigenvector Decomposition Analysis (LEiDA, Cabral et al., 2017) focuses specifically on phase-coherence of the BOLD signals, where the phases are obtained using a Hilbert Transform (Deco and Kringelbach, 2016; Deco et al., 2017; Glerean et al., 2012). Identifying TVFC states is done according to the BOLD phase relative to the leading eigenvector (i.e., largest magnitude), which captures the dominant pattern of functional connectivity per time point. Focusing on the leading eigenvector per time point deals with the main objection raised against time-frequency approaches: that the output is very extensive, which limits interpretability (Hutchison et al., 2013). Point-process and co-activation approaches. Point-process analysis assumes that all relevant information about TVFC can be extracted from short periods where BOLD signals exceed a certain threshold (Petridou et al., 2013; Tagliazucchi et al., 2016). The metric of interest here is the number of co-activations between voxels, defined by simultaneously crossing the threshold. The selection of this threshold, however, is arbitrary. Liu and Duyn (2013) suggested to only use frames where a seed time-course exceeded the threshold. Moreover, where the metric in the original method was binary (exceeded vs. not exceeded), they suggested clustering the original fMRI volumes obtained at those frames generating so-called co-activation patterns. This brings us to a second important differentiation between data-driven TVFC methods (in addition to the time frame considered), namely, whether the temporal ordering of the data is considered. Some methods ignore temporal structure altogether, for example, clustering applied to obtain co-activation patterns, which treats time points interchangeably. Other TVFC approaches make use of the temporal structure at some stages of the analysis, for example, in estimating the phase-coherence of BOLD in LEiDA, but ignore it at other stages, for example, by subsequently applying k-means clustering (Cabral et al., 2017; Lurie et al., 2020). Modelling approaches. Another line of TVFC methods aims not just to estimate time-varying functional connectivity fluctuations over time, but to explicitly model to temporal structure (Preti et al., 2017). Promising approaches of this sort fit multivariate dynamic models, that is, Hidden Markov Models (HMMs; Vidaurre et al., 2016, 2017) or autoregressive models (AR-1; Rogers et al., 2010) to fMRI time series. The most crucial difference being that HMMs assume the existence of discrete brain states, whereas AR-1 models are continuous, such that each time point is modelled as a linear combination of the previous ones. The latter approach was recently shown to more adequately capture task-based phenotypes than time-averaged functional connectivity measures (Liégeois et al., 2019), illustrating the potential added value of TVFC over time-averaged approaches. Given the high dimensionality of TVFC data, previously discussed dimensionality reductions are performed in the temporal domain (resulting in loss of information on the temporal ordering of the data). Most methods also start with a dimensionality reduction in the spatial domain, called a parcellation. Instead of analysing the temporal properties based on voxel time series directly, the voxels are first summarized at a lower dimensionality. That is, voxels are grouped into brain regions that are functionally or anatomically similar. This procedure typically results in a set of entirely separate brain regions, that is, there is no overlap between them (called a "hard parcellation"), or in regions that have minimal overlap (called a "soft parcellation"). For functional parcellations, this implies that a one-to-one mapping between cortical structure and brain function is assumed. In other words, a brain region is expected to be recruited by one functional entity only. There is no (or minimal) spatial overlap. Accounting for Spatial Overlap: TFM Analysis The lack of accounting for spatial overlap was formulated by Friston (1998) in a critique on spatial ICA. He argued that by inducing orthogonality in the spatial domain, it is assumed that functional networks do not share underlying anatomical structure. However, such spatial overlap is expected: first, because functional sub-units within certain regions are not separable due to the limited spatial resolution of fMRI data, and second, because certain areas supposedly participate in more than one functional network. In response, it was posited that a similar limitation occurs when enforcing orthogonality in the temporal domain due to stimulus-correlated effects (e.g., motion related to the presented stimuli) and that spatial ICA provides relatively adequate representations when spatial overlap occurs in the presence of noise (which is inherent to fMRI data; Beckmann et al., 2005). The amount of spatial overlap that spatial ICA accounts for is, however, limited (Bijsterbosch, 2017). Not accounting for spatial overlap can obscure the actual functional organization of the brain, in the words of Smith et al. (2012) because: "if two regions participate in multiple functional networks, their apparent temporal correlation will reflect the combined contribution from all networks, obscuring the true underlying functional organization. The correlation will not necessarily be meaningful, being some unknown combination of correlations caused by various distinct processes" (p. 3131). Smith et al. (2012) therefore addressed this by subsequently applying a spatial ICA and a temporal ICA decomposition. Time Rseries of resting-state networks are hence further decomposed (‘unmixed’) into a set of temporally independent components called "Temporal Functional Modes" (TFMs). Employing TFM analysis, it was found that well-established resting-state networks, for example, the DMN, involve different functional sub-processes that overlap spatially (Smith et al., 2012). This approach has hitherto not widely been used because it requires an enormous number of samples, not generally available in fMRI studies. Recently, TFMs were obtained at the single-subject level for the first time using an ultra-fast imaging sequence (Gomez et al., 2020). This provides the opportunity to examine whether inter-individual differences in TFMs relate to psychiatric and behavioural measures. Gomez et al. (2020) performed TFM analysis both on resting-state data and on task-based data of participants performing a visuomotor association task. They found that reproducible TFMs can be found at the individual subject level. Similar TFMs were identified independent of the original spatial dimensionality reduction, and they were deemed largely free from confound contamination. Six TFMs were identified across most participants and scans, the most common of which consisted of an anti-correlated pattern between the DMN and task-positive networks termed the "default temporal mode". TV-TFM Analysis The extent to which TFMs recruit spatial parcels (described by the TFM mixing matrix) remains time-invariant in the original TFM model. To account for the time-varying nature of functional connectivity and simultaneously account for spatial overlap, we developed a novel, time-varying version of classical TFM (TV-TFM) analysis. We show analytically that the fixed mixing matrix in classical TFM analysis is the temporal average of a time-varying mixing matrix, which can be obtained in closed form and captures moment-by-moment estimations of brain network reconfigurations. In other words, how spatial parcels are reconfigured and combined by independent processes of interest. We apply the novel method to a high-temporal resolution fMRI dataset of participants performing a visuomotor association task and demonstrate that the retrieved network reconfigurations closely follow the expectation based on the task design.
Methods[a][b] [back to top]
TV-TFM analysis pipeline
In the current study, we extended classical TFM analysis to obtain moment-by-moment estimations of brain network reconfigurations while simultaneously accounting for spatial overlap. Traditional TFM analysis consists of two stages: a spatial decomposition to retrieve canonical resting-state networks and a subsequent temporal ICA to retrieve TFMs (see panel A of Figure 1 for a visual representation). TFM stage 1: partial decomposition. As previously described, a popular tool for the first stage is spatial ICA. To identify a set of spatial building blocks, pre-processed fMRI data with v voxels measured at t time points are decomposed into a set of statistically independent components (d). That is, a two-dimensional (v × t) fMRI data matrix () is approximated by a set of spatially independent non-Gaussian sources, which are described as spatial maps (, of dimensions: v × d) and their associated time courses (, of dimensions: d × t): (1) where denotes the residual error of the spatial decomposition. Obtaining resting-state networks using spatial ICA can be done in different ways. First, spatial ICA can be applied at the single-subject level, which results in a description of subject-specific functional connectivity networks. Applying ICA at the single-subject level is problematic when the goal is to compare across participants because of the correspondence problem. That is, slight variations in the data can cause differential splitting of components, such that one component in one participant can be split into multiple components for another participant. This precludes matching of components across different datasets. A group-level ICA followed by the dual-regression technique can be performed to overcome this (Beckmann et al., 2009; Nickerson et al., 2017). Participant-specific maps and time series are obtained here through two multiple regression analyses within a general linear model (GLM) framework. First, individual subject’s time series are estimated by regressing the group ICA network template onto the individual participants’ pre-processed fMRI data (i.e., spatial regression): (2) with: (3) where represents the average time-course related to a specific network component of the template () after considering the contributions of the other components to that time series, denote the residual errors of this multiple regression analysis, and is the matrix pseudoinverse. The subject-specific time courses estimated in equation (3) are then used as predictors in a second multiple, temporal regression to estimate the subject-specific spatial maps (): (4) with: (5) where denote the residual errors of this multiple temporal regression analysis. With group-ICA and dual-regression, individual subject maps and time series are estimated with respect to group or sample-specific connectivity networks. Instead of group-specific templates, previously established functional (or anatomical) templates can also be used (Nickerson et al., 2017). This approach is taken in the current study: we performed dual-regression analysis using templates of major brain networks identified in resting-state using group spatial ICA, which were previously shown to correspond to networks identified from a large set of task-based brain activation studies (Smith et al., 2009). The latter finding provides support for the robust occurrence of these spatial networks, both in rest and during task. For constructing the templates, group-ICA was performed at a dimensionality of 20 to examine large-scale brain networks (SMITH20 template) and a higher dimensionality of 70 (SMITH70 template) to probe a lower level of the functional hierarchy (i.e., subnetworks). Of the 20 components identified with the previous, ten were well-defined and clearly overlapping the networks identified in task-based data. As previously described, Gomez et al. (2020) found that similar TFMs can be extracted across different functional (or anatomical) spatial decompositions. The SMITH20 spatial template was hence chosen for illustrative purposes in the rest of this paper because the included networks are better defined, which aids interpretability. TFM stage 2: temporal decomposition. In classical TFM analysis, the second stage consists of applying temporal ICA to the time series retrieved in the first step, decomposing them into a set of statistically independent TFMs (k). That is, the spatial ICA time series (, of dimension d × t) are approximated by a set of temporally independent non-Gaussian sources, which are described as a product of the TFM mixing matrix (, of dimensions: d × k) and the associated temporally independent time series (, of dimensions: k × t): (6) where denote the residual errors of the temporal ICA. Classical TFM analysis thus describes the pre-processed fMRI data as a product of three matrices: the spatial ICA spatial maps, the TFM mixing matrix (which describes how the temporal components weigh onto the spatial components), and the associated TFM time courses: (7) where denotes the total residual error. Temporal ICA at stage two was performed with a dimensionality of 21 in the original Smith et al. (2012) paper and at dimensionalities 8, 15, and 21 at the single-subject level by Gomez et al. (2020). Ensuring a lower dimensionality for the temporal than for the spatial decomposition (i.e., smaller than the spatial ICA matrix rank), in the current study we ran temporal ICA at dimensionalities 8, 10 and 15. The main results described in the current paper stem from the highest model order (i.e., 15). TV-TFM analysis: obtain time-varying temporal functional modes. As described in equations (6) and (7), in classical TFM analysis a time-averaged mixing matrix is obtained. In other words, the extent to which spatial parcels (e.g., canonical functional connectivity networks) are reconfigured and combined by independent processes of interest is assumed to be constant. In the current study, classical TFM analysis is extended to include moment-by-moment estimations of the extent to which TFMs reconfigure and combine different spatial components. That is, a time-varying TFM mixing matrix is obtained (see panel B of Figure 1 for a visual representation).
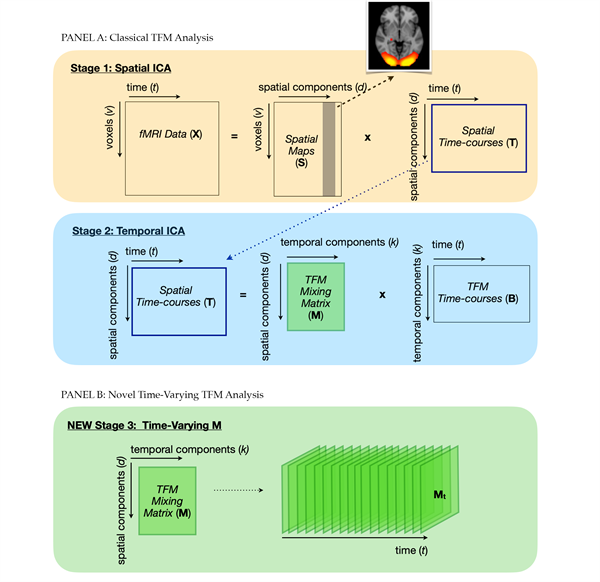
Consider the TFM model of pre-processed fMRI data as described in equation 7, where and were previously defined as the spatial maps, the TFM mixing matrix and the TFM time series, respectively. If: (8) then the TFM mixing matrix M can be described as: (9) where and the product of and amounts to the right pseudo-inverse of the TFM time courses. Note that the only time-dependent quantities in equation (9) are and . Considering the product between them in equation (9) as a sum over time: (10) where , and: (11) can be obtained by summing over the time dimension of this time dependent quantity, namely: (12) Note that for normalised (mean zero and unit standard deviation) and time series, describes a scaled version (through , i.e., the covariance of ) of the instantaneous correlation (van Oort et al., 2018) between and at time . That is, the instantaneous correlation between the spatial maps regressed onto the pre-processed fMRI data (i.e., ) and the TFM time series (). Defining a function: (13) we have that: (14) As shown in equation 14, the function is a scaled version of such that is the time-average of the instantaneous correlation between and scaled by its covariance. A python package for running TV-TFM analysis was developed. The package will be made available on https://github.com/tamarajedidja/tv-tfm upon publication. Task-based fMRI TV-TFM analysis was applied to high-temporal resolution fMRI data of participants performing three runs of a visuomotor association task, previously collected by Z. Fazal. Data were collected at the Robarts Research Institute at the University of Western Ontario in Canada. Healthy volunteers were recruited according to the guidelines of the Health Sciences Research Ethics Board of Western University. All volunteers provided written informed consent. Data were organised according to the Brain Imaging Data Structure (BIDS; Gorgolewski et al., 2016).
Sample
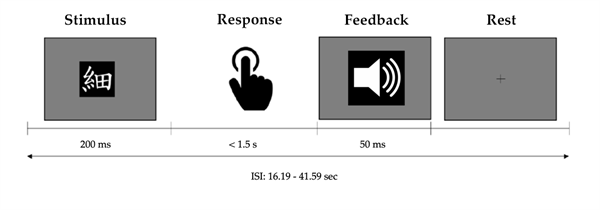
Seventeen healthy volunteers participated in this study. Inclusion criteria were right-handedness and age below 50 years old. Visual inspection of data quality resulted in the exclusion of three participants due to an acquisition error, that is, the scanner timing was not well-matched to the presentation of stimuli. This resulted in a final sample of 14 participants between 19 and 32 years old, of which seven were female (age: M = 23.43, SD = 3.36) and seven were male (age: M = 24.57, SD = 4.28). For two of the participants, one individual task run was excluded from the analysis based on insufficient brain coverage. Procedure. Participants underwent one scanning session of about 40 minutes. It consisted of three runs of a visuomotor association task while functional magnetic imaging data were acquired, and an anatomical scan at the end of the task run. Participants were trained on the visuomotor association task by performing one run before the scanning session. As visually presented in Figure 2, the task consisted of learning arbitrary associations between eight Japanese Kanji characters and four different motor responses, which were learned by trial and error. Between 25 and 36 (M = 31.14, SD = 1.68) visual stimuli were presented for 200ms. Participants were required to respond as soon as possible, but at least within 1.5s after stimulus onset. Motor responses consisted of pressing one of four buttons on a fibre optic response pad, where each button corresponded to two different Kanji characters. The response pad was placed on the abdomen of the participant, who consistently kept the index, middle, ring, and little fingers of the right hand on corresponding buttons. Performance feedback was given immediately after the response using three different sounds indicating a correct, incorrect, or too late response, respectively. Feedback was presented for 50ms, after which a fixation cross appeared until the next trial. Long inter-stimulus intervals (ISI) ranging between 16.19 and 41.59s (one outlier of 80.47s; M = 19.46, SD = 3.28) were used, to ensure that the haemodynamic response would return to baseline before the next trial. Presentation of visual stimuli and registering motor responses was done using Presentation 20.1 (Neurobehavioral Systems, San Francisco, CA).
Data acquisition
Data were acquired on a Siemens MAGNETOM 7T MR scanner, equipped with a 32-channel head coil. The use of a highly optimized AC84 (GenII) head gradient engine with a maximum gradient strength of 80 mT/m and 2nd, 3rd, and 4th order shims allowed the acquisition of T2* images at a very high temporal resolution. Functional images were acquired using a 2D multiband gradient-echo EPI protocol with an interleaved slice acquisition sequence (TR = 206ms, TE = 22ms, voxel sizes = 2.70 x 2.70 x 3.11 mm, flip angle = 20°, matrix size = 86 x 86, 32 slices, slice thickness/gap = 2.7 mm/3.1 mm, acceleration factor = 8). A total of 3000 volumes were obtained per task run of about 10 minutes, amounting to 9000 task-based volumes across task runs, per participant. Image reconstruction was performed using Slice-GRAPPA (Cauley et al., 2014; Hoge et al., 2018; Setsompop et al., 2012). For registration purposes, anatomical images were acquired using the MP2RAGE sequence (Marques et al., 2010; TR = 6s, TE = 2.3ms, voxel sizes = 1 mm isotropic, flip angles = 4 and 5 degrees; TI1 = 0.8s, TI2 = 2.7s; matrix = 240 x 240; 240 slices of 1 mm slice thickness).
Data pre-processing
Data were pre-processed using fMRIPREP version 1.1.6 (Esteban et al., 2019), which is a Nipype-based tool (version 1.1.2; Gorgolewski, et al. 2011; Gorgolewski et al., 2018). Anatomical scans were corrected for spatial intensity variations using N4BiasFieldCorrection (ANTs 2.2.0; Tustison et al., 2010). A T1-weighted reference map was computed after registering the T1-weighted images using the mri_robust_template (Freesurfer, 6.0.0; Reuter et al., 2010). The T1-weighted reference was brain extracted using antsBrainExtraction.sh (ANTs 2.2.0) with a target template from the Open Access Series of Imaging Studies. Nonlinear registration to the ICBM 152 Nonlinear Asymmetrical template version 2009c (Fonov et al., 2009) was performed using antsRegistration (ANTs 2.2.0; Avants et al., 2008). The pre-processed T1-weighted reference map was used as a reference for functional registration. Brain-tissue segmentation of the T1-weighted image into cerebrospinal fluid (CSF), white matter (WM), and grey matter (GM) was performed on the brain-extracted T1w using FAST (FSL v.6.0.0; Zhang et al., 2001). For all three task-based fMRI runs per participant, the following pre-processing steps were taken. Motion correction parameters were calculated using MCFLIRT (FSL 6.0.0, Jenkinson et al. 2002). A reference volume and its brain extracted version were generated using a custom methodology of fMRIPREP, that is, averaging non-steady state volumes and computing a brain mask with init_enhance_and_skullstrip_bold_wf. Susceptibility distortions were estimated using a field map-less approach, that is, registering the functional reference image to the T1w reference using intensity inversion (Huntenberg, 2014; Wang et al., 2017) while regularising by constraining deformation to be nonzero only along the phase-encoding direction and modulation with an average field map template (Treiber et al., 2016) as implemented in ANTs 2.2.0. The transformation parameters for co-registration to the T1w reference were calculated with MCFLIRT (FSL 6.0.0, Jenkinson et al., 2002), using boundary-based registration (Greve and Fischl, 2009) with 9 degrees of freedom. A concatenated transform to correct for head-motion, susceptibility distortion, functional to T1w registration, and registration from T1w to MNI space (i.e., MNI152NLin2009cAsym) was applied in one step using Lanczos interpolation (Lanczos, 1964), as implemented in antsApplyTransforms. Following Gomez et al. (2020) several confounding time series were computed, which were not used to denoise the functional time series but to measure whether they correlate to the TFM time series. This concerned: three region-wise global signals, extracted within the CSF, the WM, and the whole-brain mask. The head-motion estimates from MCFLIRT were also placed within the corresponding confounds file. Datasets were temporally high-pass filtered with a cut-off frequency of 0.01 Hz (i.e., a periodicity of 100s) and spatially smoothed with a FWHM of 5.4mm. ICA-based denoising of the data was performed. Each pre-processed run was decomposed into 70 spatially independent components using Probabilistic Independent Component Analysis (Beckmann & Smith, 2004) as implemented in MELODIC (version 3.15, FSL). The following data pre-processing was applied to the input data: masking of non-brain voxels, voxel-wise de-meaning of the data, normalisation of the voxel-wise variance, and subsequent whitening and projection into a 70-dimensional subspace using Principal Component Analysis. The whitened observations were decomposed using spatial ICA (Hyvarinen, 1999). Estimated component maps were divided by the standard deviation of the residual noise and thresholded by fitting a mixture model to the histogram of intensity values (Beckmann & Smith, 2004). IC components were manually classified into signal and noise, and the pre-processed data were denoised using FSL REGFILT (FSL v.6.0.0, Jenkinson et al., 2012). The pre-processed data were moreover variance normalised (mean zero and unit standard deviation across time) using FSLMATHs (FSL v.6.0.0, Jenkinson et al., 2012).
Statistics
Explorative statistical analyses were conducted at the single task-run, single-subject and at the group level. The well-established dual-regression technique (stage 1), the classical time-averaged TFM model (stage 2) and the new TV-TFM model (new stage 3) were exploratively compared in their ability to capture task-based variation in high temporal resolution fMRI data of participants performing a visuomotor association task. Single-subject results are illustratively presented for one task run of one participant (subsequently referred to as ’the selected dataset’). Criteria for selecting this dataset for illustrative purposes are outlined in section II of the Supplementary Material. First, it was examined how well task-related variability is captured: i) in the dual-regression time series; ii) in the TFM time series as obtained with classical TFM analysis, and iii) in the time-resolved mapping of TFMs onto the resting-state networks as obtained with TV-TFM analysis. Therefore, at each stage of the analysis pipeline (dual-regression, classical TFM analysis, and TV-TFM analysis), a first-level analysis was run separately for each task run, consisting of pairwise Pearson correlations between the time series and the standardised task regressors of the visuomotor association task (i.e., visual stimulation and motor response). To account for dependencies between task runs per participant, for stage one (dual-regression), a fixed-effects analysis was run on the Fisher-Z transformed correlation values of the association between the dual-regression time series and the task regressors. That is, they were averaged over task-runs, per participant. To identify the resting-state network time series that across participants were significantly related to the task, a third-level analysis was performed using a one-sample t-test per network. At stage two of the analysis pipeline (classical TFM analysis), group-level comparisons per TFM are problematic due to the correspondence problem of temporal ICA. Therefore, it was examined whether per task-run TFM time series (i.e., B) better capture task-related variability than dual-regression time series (i.e., T). This was done by first selecting the dual-regression time series and the TFM time series that showed the strongest absolute correlation to the visual task regressor. The Fisher-Z transforms of these absolute correlation values were then compared per task-run using a paired sample t-test. A proof-of-concept for TV-TFM analysis was obtained using trial-averaging, where time-varying mixing matrix network weights (a proxy for network involvement) were averaged over trials. The DMN is widely recognised as a task-negative network (Singh and Fawcett, 2008). Therefore, we tested whether we observed an unfolding, increasing difference between the DMN and task-positive network (i.e. visual, sensorimotor and auditory) weights in entries of the time-varying mixing matrix, for task-related TFMs. It was first examined whether this pattern was observed across trials. Therefore, per task run, entries of the time-varying mixing matrix that corresponded to the TFM that showed the strongest absolute correlation with the visual task regressor were averaged across trials. A trial consisted of a 60-frame epoch (i.e., 60 × 206ms = 12.36s), starting at the onset of the visual stimulus. This interval was chosen such that it covers most of the expected time it takes for the haemodynamic response to increase in response to the visual stimulus and planning a motor response as well as come back to baseline, while not overlapping between trials. Note that if the strongest correlation was negative, the direction of the time series was reversed because it is not informative (i.e., it depends on the sign of the mixing matrix weight). Differences between the trial-averaged DMN weight and each of the three trial-averaged task-positive network weights (i.e., the average of the primary and lateral visual weights, the sensorimotor weight and the auditory weight) were then separately tested per time frame using a paired t-test. To account for dependencies between task runs per participant, a fixed-effects analysis was run. That is, the t-values were averaged across task runs per participant. Lastly, it was examined whether an increasing difference between the DMN weight and each of the task-positive network weights in response to the task was observed across participants. This was done by computing a group-level Z-value per time point, for the comparison between the DMN and each of the task-positive networks, separately.
Results [back to top]
Stage 1: Spatial decomposition Dual-regression with respect to the brain network templates by Smith et al. (2009), resulted in a set of canonical resting-state networks per task run, per subject, and per spatial network template (SMITH20 and SMITH70 templates). As previously mentioned, the results stem from a spatial decomposition using dual-regression against the SMITH20 template. The resulting resting-state networks are described by a spatial map (see, e.g., Figure 3) and a corresponding time series (see, e.g., panel A of Figure 4).

Bonferroni corrected, Pairwise Pearson correlations between the dual-regression time series and the visuomotor association task regressors (i.e., visual stimulation and motor response) are illustratively presented for the selected dataset in panel B of Figure 4. Whether the group averages of the Fisher-Z transformed correlation values were significantly different from zero was tested for each network with a one-sample t-test. Bonferroni corrected results are presented in panel C of Figure 4.
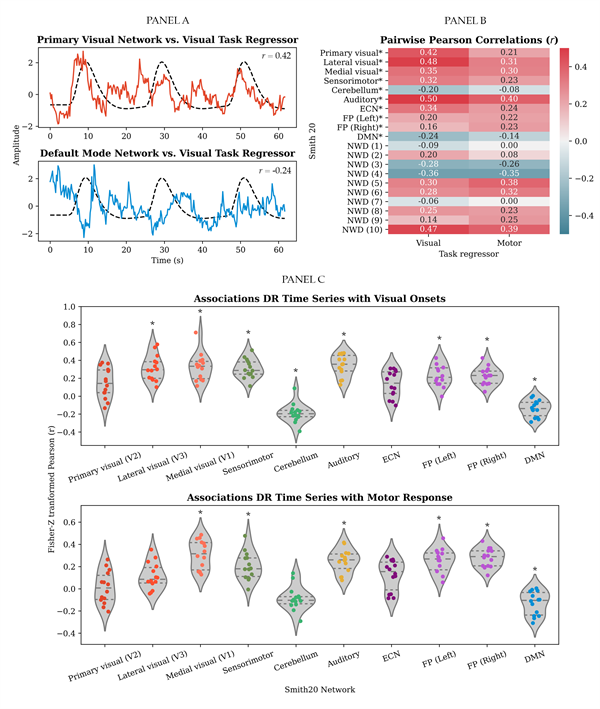
The following findings are relevant for the purpose of the present paper. As visually represented in Figure 4, across participants correlations with the task regressors were found across most of the networks. This indicates that task-related information is distributed across the dual-regression time series of the different resting-state networks. Note that contrary to the expectation, the primary visual network (V2) did not significantly relate to the onsets of the visual stimulus, Fisher-Z = 0.14, t(13) = 3.25, p = .006 (after Bonferroni correction; i.e., significance below .05/20 = .0025). Please refer to the Supplementary Material, section III for an overview of the statistics of all one-sample t-tests. Stage 2: Temporal decomposition Temporal ICA resulted in a set of TFMs per task run, per subject, per spatial network template (SMITH20 and 70) and per temporal ICA dimensionality (8, 10, 15, and 21 depending on the model order of the spatial template). As previously addressed, the results stem from the highest model order, that is, 15, for the SMITH20 template. TFMs are described in terms of their weighting onto the SMITH templates (node weights/a mixing matrix column, see Figure 5) and the corresponding time series. TFM time series were related to confound regressors as an indication of their neuronal basis. Results are presented in section IV of the Supplementary Material. As visually illustrated for the selected dataset in panel A of Figure 5, we identified a "task-positive mode" which includes regions related to the visual, motor and auditory aspects of the task, that is, primary visual (occipital pole), medial visual (e.g., lingual and occipital fusiform gyri) and lateral occipital regions; the temporal pole; motor regions along the dorsal stream (e.g., primary motor cortex and supplementary motor cortex) and primary auditory cortex. A similar component was identified across most task runs and participants. Please refer to section V of the Supplementary Material for further details. We identified a similar component to the most common TFM identified by Gomez et al. (2020), the "default temporal mode", consisting of core hubs of the DMN (i.e., angular gyrus and precuneus) in anti-correlation with the motor regions (superior parietal cortex, post-central gyrus and supplementary motor cortex), the auditory network, and the insula (panel B of Figure 5). A similar component was identified across most task runs and participants. Please refer to section V of the Supplementary Material for further details. Note that contrary to the findings by Gomez et al. (2020) no anti-correlations with visual areas were found for the selected dataset. This potentially results from component splitting, since a temporal ICA of model-order ten performed on the time series of the ten well-defined networks of the SMITH20 template identified a component similar to the default temporal mode also in anti-correlation to visual areas (see the Supplementary Material section V).
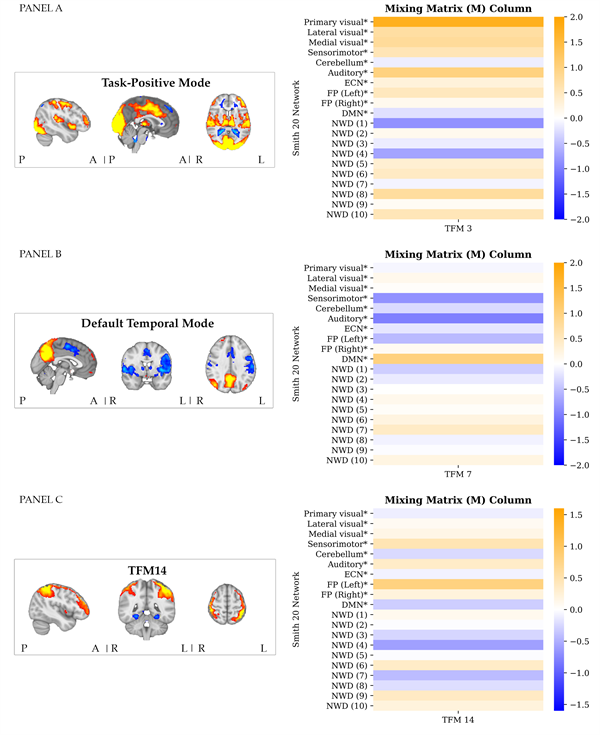
Bonferroni corrected, Pairwise Pearson correlations between the TFM time series and the visuomotor association task regressors are illustratively presented for the selected dataset in panel A of Figure 6. The previously described "task-positive mode" (TFM3) correlated most strongly with the visual task regressor. TFM14 correlated most strongly with the motor task regressor. A visual representation of the spatial map of TFM14 is presented in panel C of Figure 5. It consists of an anti-correlated pattern between the frontoparietal attention network and the temporal fusiform cortex, a higher-level visual processing area involved in sensory integration and memory. In panel B of Figure 6, correlation values are plotted per task run for the associations between the time series of the TFMs that correlated most strongly with the visual (left) and motor (right) task-regressor and those regressors, respectively.
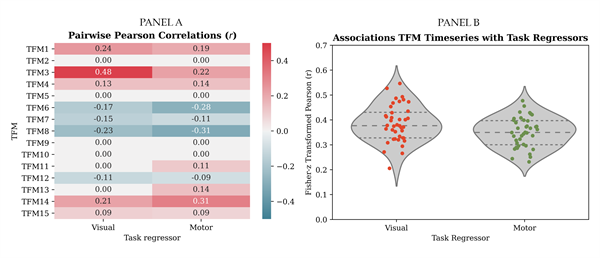
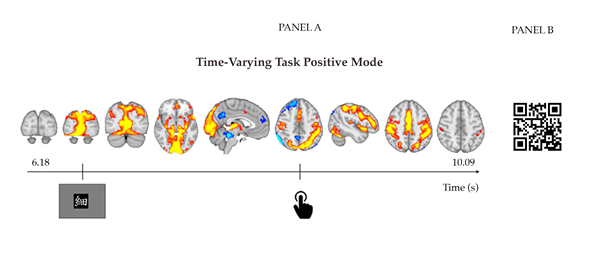
A relevant observation for the purpose of the present paper is the increased sparseness of the correlation matrix as observed in panel A of Figure 6, compared to the dual-regression correlation matrix in panel B of Figure 4. That is, task-related variation appears to be represented in fewer components. This corresponds to the fact that temporal ICA retrieves temporally independent processes, consisting of linear combinations of resting-state networks. Moreover, it was found that across task runs, absolute correlations with the visual task regressor were stronger for TFMs (which per task run correlated most strongly to the task) than for dual-regression time series (which per task run correlated most strongly to the task), paired-t(39) = 6.38, p < .0001. This is in line with the conceptual expectation that task-related processes are temporally independent of other brain processes and can be captured using temporal ICA. New stage: Time-varying temporal functional modes TV-TFM analysis resulted in time-varying mixing matrices (i.e., Mt and its scaled version f ) per i) task run, ii) participant, iii) spatial network template (SMITH 20 and 70), and: iv) temporal ICA dimensionality (8, 10, 15, and 21 depending on the spatial model order). As previously described, the results stem from the highest model order (15) for the better defined SMITH20 template. TV-TFMs can be described spatially by means of the product between the spatial maps, S, and the entries of the time-varying mixing matrix related to one TFM component. This allows per time point visualisations of the resting-state networks (as identified by Smith et al., 2009) recruited by this TFM. An example for the selected dataset is presented in Figure 7, where the temporal evolvement of the task-positive mode (TFM3) is presented for one trial. Onsets of the visual stimulus and motor response are presented at the bottom of the Figure and are operationalised as the first time frame that the stimulus/response convolved with the haemodynamic response function exceeds zero. Time-locked to the visual stimulus, we observe recruitment of primary visual areas, followed by medial and lateral visual areas. Time-locked to the motor response, we observe the recruitment of superior parietal motor regions and subsequent recruitment of the pre-central gyrus/primary motor cortex.
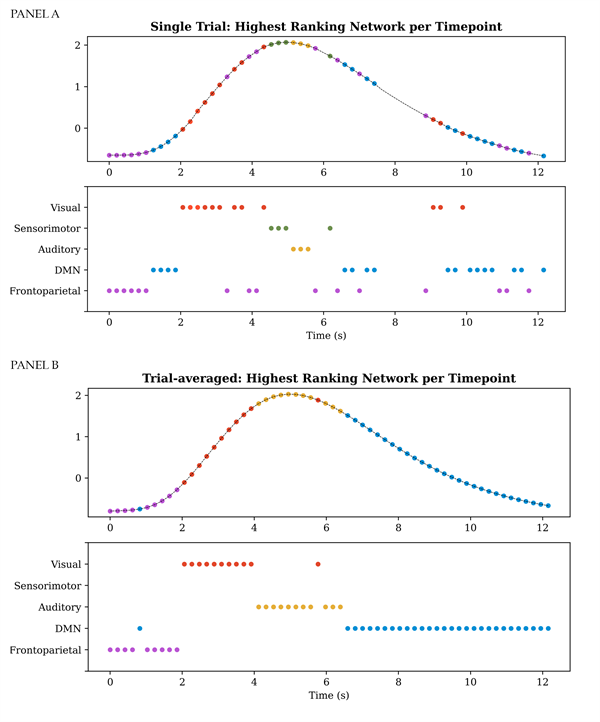
Ranking-ordering the entries of the time-varying mixing matrix per TFM, per time-point, allows representing which networks are most strongly recruited by the respective TFM over time. A single trial example for the task-positive mode (TFM3) is presented in panel A of Figure 8 for the selected dataset. The bottom figure in panel A shows per network when in time its weighting was the strongest (i.e., it had the highest rank). The top figure in panel A shows the highest-ranking network per time point plotted on the visual task regressor. We observe that the ranks closely follow the expectation based on the task design. Before the onset of the stimulus the frontoparietal attention and default mode networks predominate. In response to the onset of the visual stimulus the visual networks predominate, followed by short periods in which the sensorimotor network and the auditory network subsequently rank highest. Averaging over all trials for the selected dataset, we observe a similar pattern except that predominance of the sensorimotor network is not observed (see panel B).
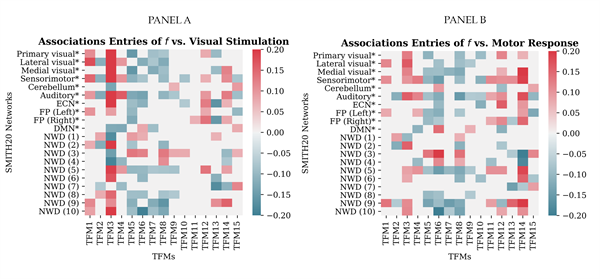
Pairwise Pearson correlations between each entry of the time-varying mixing matrix and the visuomotor-association task regressors are illustratively presented for the selected dataset in Figure 9. The sum of the correlation values across the entries of each TFM was highest for the task-positive mode (TFM3) for the visual task regressor and for TFM14 for the motor task regressor (see Supplementary Material, section VI). These are the same TFMs for which the time series were most strongly related to the task regressors (as concluded in stage two of the analysis pipeline; see panel A of Figure 6). In line with the expectation, correlations between the visual task regressor and the TFM3 entries that correspond to the task-positive networks (i.e., primary and lateral visual, sensorimotor and auditory) are opposite to the correlation between the visual task regressor and the TFM3 entry that corresponds to the DMN. This pattern is slightly different for the correlations between the motor task regressor and the entries of TFM14, in the sense that no significant correlations with the lateral visual network (V3) and the DMN were found.
The event-related approach, consisting of trial-averaging the time-varying mixing matrix, provided the main proof-of-concept for TV-TFM analysis. Panel A and B of Figure 10 show single-subject results for the selected dataset. Trial-averaged entries of the time-varying mixing matrix for the task-positive mode closely follow the expectation based on the task design, that is, starting with an increase in the primary visual network, followed by increases in the sensorimotor network and the auditory network, and a concurrent decrease in the DMN weights (see panel A). For comparison, in panel B, an example is plotted for a TFM for which the time series were not significantly related to the task. A similar pattern was found across most task runs and participants. Please refer to section VII of the Supplementary Material for trial-averaged plots of task-relevant entries of the time-varying mixing matrix per task run, per participant. Support for the observed trend was obtained using per time frame paired t-tests on the differences between the DMN entry and each task-relevant entry of the time-varying mixing matrix separately. That is, DMN versus the average of the primary and lateral visual weights, DMN versus the sensorimotor weight and DMN versus the auditory weight. Group-level results (after averaging across task runs per participant at the second level) for the TFMs that showed the strongest absolute correlation with the visual task regressor are presented in panel C of Figure 10. Across trials, task runs and participants, the differences between the DMN and: i) the average of the primary and lateral visual entries, ii) the sensorimotor entry, and iii) the auditory entry increased subsequently.
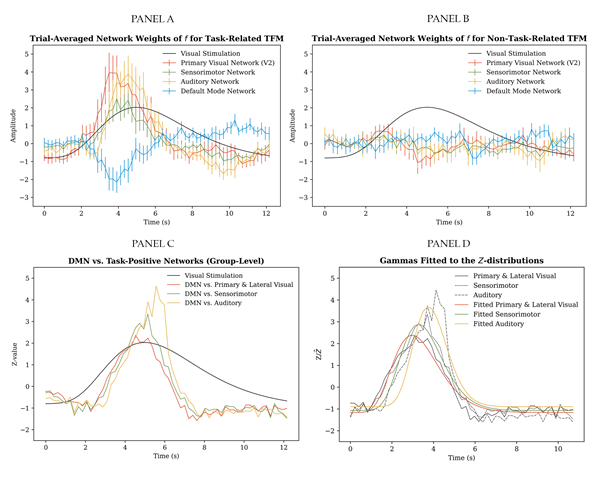
As a first parametrisation, the Z-distributions in panel C of Figure 10 were fitted with three independent gamma distributions, with: (15) Model fitting was performed after restricting the time range, shifting the distributions up into the positive domain of Z and normalising the distributions (such that they integrate to one). The parameters for these operations are presented on the left side of Table 1. The parameters for the best fitting gamma distributions are presented on the right side of Table 1. These distributional parameters reaffirm subsequent recruitment of the visual, sensorimotor and auditory networks by task-related TFMs (see panel D of Figure 10).
| Operation Parameters* | Distributional Parameters** | |||||
| time range (s) | shift in y |
α |
β |
Mode (s) | ||
| Visual | 1.44 - 12.36 | 1.16 | 8.37 | 0.51 | 2.98 | |
| Sensorimotor | 1.44 = 12.36 | 1.07 | 11.82 | 0.68 | 3.28 | |
| Auditory | 1.44 - 12.36 | 0.90 | 22.47 | 1.18 | 3.75 | |
(*) Operation parameters regard the preparatory steps before model fitting (i.e., restricting the time range, shifting the distributions into the positive domain).
(**) Distributional parameters for the optimal (least squared error) model fit are presented.
Discussion [back to top]
Proof-of-Concept
With the current study, we provided the first proof-of-concept for TV-TFM analysis. The goal was to develop a TVFC method that accounts for spatial overlap. The latter is inherently safeguarded by extending classical TFM analysis, which accounts for spatial overlap by following a spatial decomposition with a subsequent temporal ICA on the retrieved time series. That different temporal processes indeed recruited the same spatial components shows from the fact that different temporally independent processes (TFMs) map onto the same spatial maps. Regarding the first, TV-TFM analysis obtains moment-by-moment estimations of the extent to which different spatial components (e.g., resting-state networks) are reconfigured by temporally independent processes (TFMs). A first validation was provided by applying TV-TFM analysis to task-based fMRI data and subsequent trial-averaging of the time-varying mixing matrix entries per resting-state network for task-related TFMs. The pattern observed across participants, sessions, and trials closely followed the expectation based on the task design. We observed an increase in visual areas, the sensorimotor network, and the auditory network subsequently, and a concurrent decrease in DMN weights. Differences in timing between these subsequent increases were minor. This is inherent to the task design, in which the different processes follow each other up in close succession: the visual stimuli are presented for 200ms, participants are required to respond as soon as possible using a button press and get immediate auditory feedback. Previous attempts to temporally disentangle these processes in high-temporal resolution fMRI data by means of a mass univariate GLM approach were unsuccessful (Noteboom, 2018). At the very short TR of .206ms in the current study, we could not disentangle the processes using dual-regression with respect to existing resting-state network templates either. In part, these processes were also considered a joint independent temporal process as identified by temporal ICA. That is, applying temporal ICA at stage two of the analysis pipeline resulted in a task-positive component that included brain regions implied in all these processes (i.e., visual, motor and auditory). TV-TFM analysis proved to be a step forward towards temporally disentangling them. Notably, event-related averaging could also have been applied to T, that is, the dual-regression time series. This would have provided a linear mix of resting-state networks at a whole brain level. However, with the approach taken in the current study, we retrieved a linear mix of temporally independent processes. We argue that this is a reasonable approach because it aids interpretability by reducing the dimensionality to task-related information (i.e., task-related TFMs). Assumptions and Limitations The latter points to an important assumption for this proof-of-concept, namely that temporal ICA at stage two of the analysis adequately identifies the process of interest. Preliminary support for this was provided by Gomez et al. (2020), who concluded that TFM time series carry an imprint of the task design. Our present results provide some additional support. First, we found that the TFM time series were more strongly associated with the task than the dual-regression time series. Secondly, we observed that classical (time-averaged) TFM analysis increases specificity in detecting task-related variability compared to the dual-regression approach. That is, correlations of dual-regression time series with the task were widespread across networks, whereas correlations to the task were identified for a smaller number of TFMs. The task-related TFM components moreover spatially involved regions expected to be involved in the visuomotor association task. However, studies probing the ability of temporal ICA, or classical TFM analysis specifically, to adequately capture task-related variability are scarce. This hence requires further assessment. In this study, we used a simple but effective approach to probe whether the identified temporally independent processes were task-related, that is, temporal correlation. This was decided, in part, because of multicollinearity among the task regressors. For the same task paradigm, this was also found by Noteboom (2018), who suggested that this might be a primary reason why the GLM cannot temporally disentangle these processes. However, for other task paradigms, we suggest using regression analyses within a general linear model, which allows adjusting for covariates such as handedness (note that in the current study this specific example was addressed in the inclusion criteria). Other, more advanced, for example, PCA-based, supervised approaches could also be explored. In this study we obtained the instantaneous correlations between the individual spatial maps regressed onto the noiseless reconstruction of the fMRI data, and the TFM time series (scaled by their covariance) to retrieve moment-by-moment estimations of brain network reconfigurations. If a process of interest is not captured adequately using classical TFM analysis, the task-relevant information is unexplained, residual variance. In that case, it could well be that the reconfigurations are better characterised using the original instead of the noiseless reconstruction of fMRI data. That is, accounting for the residual errors should hence be considered in the future. Future Directions We see several potential courses of action to improve and validate the TV-TFM method further. First of all, the Z-values describing the differences between the network entries of the time-varying mixing matrix across trials, task runs, and participants were not statistically tested for the current proof-of-concept. Note that doing so in the future requires non-parametric permutation testing since the averaging of t-values at the second level breaks the normality assumption (i.e., it is t–, not normally distributed). In this study, we dealt with the large amount of high-frequency content that characterises the time-varying mixing matrices by averaging over trials. In the future, we would like to apply this method at the single-trial level, which likely requires smoothing of the time-varying mixing matrix, using a Kalman filter, for example. In this study, we parametrised the group-level Z-distributions by fitting independent Gamma distributions. We aim to fit a mixture-model (potentially of another distribution, for example, Inverse Gaussian) to the Z-distributions at the single-trial level in the future, followed by a PCA analysis. Herewith, differences in timing or shape of these distributions would potentially allow predicting behavioural measures such as success vs unsuccessful trials or attentiveness vs inattentiveness to the task. Another important follow-up step is developing a dual-regression-like approach to deal with the correspondence problem at stage two, temporal ICA, to probe and compare similar temporally independent processes of interest across participants. Moreover, although Gomez et al. (2020) found that similar TFM components are found independent of the initial spatial dimensionality reduction, we suggest that it might be relevant to explore the results of TV-TFM analysis after different initial spatial dimensionality reductions. For example, the finer-grained SMITH70 template. The SMITH20 template used in the current study was obtained with a model order that has proven to robustly identify the canonical large-scale resting-state networks in resting-state fMRI (Smith et al., 2009). A model order of 20 was also shown to be optimal for decompositions of BrainMap-based ICA components. At higher granularities, subnetworks at lower levels of the functional hierarchy are identified (Ray et al., 2013). This level of detail would potentially especially be of interest for task-based fMRI data, given the idiosyncrasies of cognitive tasks. Brain network reconfigurations as obtained with TV-TFM analysis after anatomical (instead of functional) parcellations could also be explored. Contribution and Potential Applications We believe TVTFM analysis makes a relevant contribution to the current landscape of TVFC methods by probing, at a high temporal resolution, how temporally independent processes of interest reconfigure and combine spatial building blocks. As previously described, the current TVFC methods can be differentiated based on the time window they consider, ranging from instantaneous measures to time windows of 30-60 seconds standard in sliding-window analyses (Preti et al., 2017). TV-TFM analysis provides moment-by-moment, or per timeframe, estimations of TVFC. It was previously addressed that several dimensionality reductions are generally performed in TVFC analysis. Instead of ignoring temporal ordering of the data, as is done in several TVFC methods by means of k-means clustering, for example, the dimensionality reduction consists of zooming in on one or more temporally independent processes of interest which are derived in a data-driven manner (using temporal ICA). We moreover showed that the novel method is applicable to task-based fMRI data. Whereas commonly spatial brain properties are examined, the novel method allows probing temporal properties of functional integration, potentially associated with cognitive function. Although arguably the complexity of the present method is relatively high (i.e., with respect to the number of analysis steps and the mathematical derivation), we believe that interpretability is a strong point because the estimated brain network reconfigurations are described in space and time for specific processes of interests. In this task-based study, processes of interest were identified by correlation to the task regressors. This could also be done in different manners, for example, based on physiological arousal measures, which would allow extending this method to resting-state fMRI. The developed model has numerous potential applications. For example, temporal ICA could be applied to a region of interest, such that the time-varying mixing matrix describes how sub-parcels interact over time, for temporal processes of interest. Instead of using dual-regression and temporal ICA, the TV-TFM analysis can also be implemented for other initial spatial parcellations and other subsequent multivariate dimensionality reduction methods. For the latter, a supervised PCA-based multivariate method like SPADE (Llera et al., 2020) would be an option, where moment-by-moment estimations are obtained for so-called filters which optimally discriminate between certain conditions. Irrespective of the application, the goal is eventually to utilise the information present in the time-varying matrix, for example, by describing the timing or shape differences of the Z-distributions in different states or conditions and between different individuals. These parameters describe inter-individual differences in brain network reconfigurations (while accounting for spatial overlap). To conclude, we hope that in the future, this new method will further elucidate the temporal properties of functional brain networks and that it will have a significant clinical contribution, for example, by probing temporal differences in brain network reconfigurations between individuals.
Conclusion [back to top]
We presented a novel method for probing time-varying functional brain network reconfigurations. It retrieves per time point estimates of the extent to which spatial parcels are reconfigured and combined by independent processes of interest. The method can hence be utilized to probe the temporal characteristics of network involvement and how these relate to cognition and behaviour. Supplementary Material 1. Data and code availability statement Data will be stored at the Donders Repository and are made available upon request. A documented python package was developed for TV-TFM analysis, which will be made available under https://github.com/tamarajedidja/tv-tfm upon publication. The package only supports python 3.6+. 2. Dataset selection A single-subject, single task run dataset was selected for illustrative purposes. This dataset was initially selected based on the numbers of spatial correlations larger than r = .3 (an arbitrary threshold) between the well-defined networks of the SMITH20 template and TFMs (see Table 2). Given the assumption for this task-based proof-of-concept that temporal ICA at stage two of the analysis needs to identify the process of interest adequately, we selected task-related TFMs based on their association with the visual task regressor per task run. Of the participants for which all three sessions were included, the correlation between the task-related TFM and the task regressor was strongest for the selected dataset (r = .48; Fisher-Z transformed correlations are visually presented for all task runs in panel B of Figure 6).
Table 2. Spatial correlations between the ten well-defined SMITH20 networks and task-related TFMs per participant.
|
Participant |
Descriptives |
|
|
Range |
No. of r > 0.3 (ses-01,02,03)* |
|
|
1 |
<0.001–0.38 |
3, 2, 4 |
|
2 |
0.001–0.41 |
2, 2, 3 |
|
3 |
<0.001–0.42 |
3, 4, 2 |
|
4 |
<0.001–0.48 |
3, 4, 4 |
|
5 |
<0.001–0.40 |
2, 0, 4 |
|
6 |
<0.001–0.31 |
0, 7, 3 |
|
7 |
<0.001–0.34 |
2, 1, 3 |
|
8 |
0.001–0.30 |
0, 1 |
|
9 |
<0.001–0.45 |
3, 4, 8 |
|
10 |
<0.001–0.34 |
2, 2, 4 |
|
11 |
0.001–0.38 |
2, 3, 4 |
|
12 |
0.001–0.44 |
2, 6 |
|
13 |
0.001–0.41 |
2, 3, 1 |
|
14 |
<0.001–0.45 |
6, 7, 2 |
(*) Number of correlations between the well-defined networks of the SMITH20 template and task-related TFMs that are larger than r = .3.
3. Stage 1 dual-regression: statistics one-sample t-tests To identify the dual-regression time series that across participants were significantly related to the task, a third-level analysis was performed using a one-sample t-test per resting-state network as identified by Smith et al. (2009). An overview of the Bonferroni corrected t-statistics is presented in Table 3.
Table 3. Group-level results: one-sample t-test on Fisher-Z transformed correlations of task vs. dual-regression time series.
|
Participant |
Descriptives |
||
|
Visual |
Motor |
||
|
t-value |
p-value t-value |
p-value |
|
|
Primary visual (V2) |
3.25 |
0.006 0.29 |
0.7768 |
|
Lateral visual (V3) |
8.30 |
< 0.0001* 3.55 |
0.0036 |
|
Medial visual (V1) |
7.95 |
< 0.0001* 8.86 |
< 0.0001* |
|
Sensorimotor |
10.76 |
< 0.0001* 6.05 |
< 0.0001*. |
|
Cerebellum |
-6.58 |
< 0.0001* -3.00 |
0.0102 |
|
Auditory |
10.61 |
< 0.0001* 8.79 |
< 0.0001* |
|
ECN |
3.43 |
0.0045 3.37 |
0.0050 |
|
Frontoparietal (Left) |
6.93 |
< 0.0001* 8.90 |
< 0.0001* |
|
Frontoparietal (Right) |
8.49 |
< 0.0001* 12.13 |
< 0.0001* |
|
DMN |
-5.46 |
0.0001* -4.58 |
0.0005* |
|
NWD (1) |
-3.93 |
0.0017* -2.62 |
0.0212 |
|
NWD (2) |
-0.77 |
0.4555 -2.22 |
0.0448 |
|
NWD (3) |
-4.65 |
0.0005* -6.86 |
< 0.0001* |
|
NWD (4) |
-6.96 |
< 0.0001* -5.38 |
0.0001* |
|
NWD (5) |
12.90 |
< 0.0001* 14.22 |
< 0.0001* |
|
NWD (6) |
3.67 |
0.0028 2.40 |
0.0318 |
|
NWD (7) |
3.74 |
0.0025 4.44 |
0.0007* |
|
NWD (8) |
0.87 |
.3987 0.59 |
0.5671 |
|
NWD (9) |
6.33 |
< 0.0001* 6.42 |
< 0.0001* |
|
NWD (10) |
7.80 |
< 0.0001* 8.17 |
< 0.0001* |
Abbreviations: ECN = executive control network, DMN = default mode network, NWD = not well defined; (*) Significantly different from zero, after Bonferroni correction (i.e. significance below .05/20 = .0025)
4. Stage 2 classical TFM: correlation to confounds TFM time series were correlated to confound time series as an indication of their neuronal basis. As described in the main manuscript, group-level comparisons per TFM are problematic due to the correspondence problem of temporal ICA. Therefore, the task-related TFMs were selected based on the strongest absolute correlation with the visual task regressor. Their time series were correlated with the confound regressors. The Fisher-Z transformed correlation values for all task runs are presented in Figure 11. Evident from this picture is the enlarged associations with the global signal. This finding is in line with the conclusion by Glasser et al. (2018), that spatial ICA (as used for data cleaning in the current study) is very effective at removing spatially specific structured noise from high temporal resolution fMRI data, but that it can not selectively and completely remove global structured noise while retaining global signal form neural activity. Clean up using temporal ICA was proposed as a potential solution, which could be considered in the future.

5. Stage 2 classical TFM: identified TFMs TFMs similar to the task-positive and default temporal mode identified for the selected dataset were found across most participants and task runs. Spatial correlations between the node weights of M for these TFMs identified for the selected dataset and the node weights of all other extracted TFMs were computed per task run. The average spatial correlation with the Task-Positive Mode wasr = 0.64 (SD = 0.10, range: 0.34 - 0.86) and the average spatial correlation the Default Temporal Mode was r = 0.61 (SD = 0.07, range: 0.49 - 0.79). In panel A of Figure 12, two illustrative spatial maps are plotted for the TFMs that correlated most strongly to the task-positive mode identified for the selected dataset. Moreover, two illustrative spatial maps are plotted for the TFMs that correlated most strongly to the default temporal mode identified for the selected dataset in panel B.
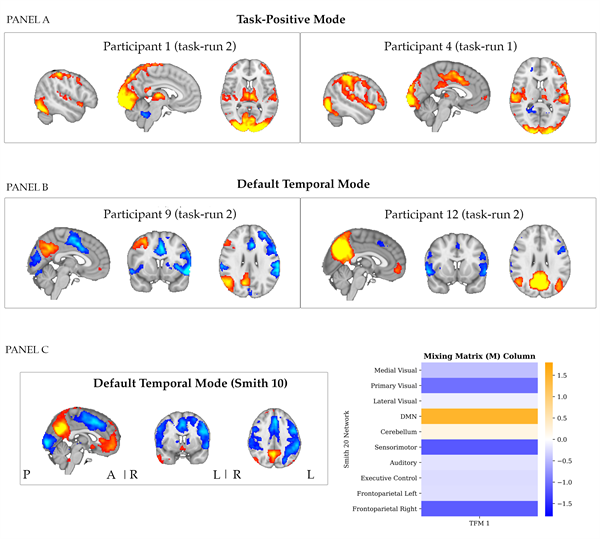
In panel C of Figure 12, the spatial map of the default temporal mode is presented for the selected dataset, as obtained with a temporal ICA of model-order ten on the time series of the ten-well defined networks by Smith et al. (2009). We observe that at this model-order, the default temporal mode involves an anti-correlated pattern between core hubs of the DMN also in anti-correlation with visual areas. 6. Stage 3 TV-TFM: correlation values for all entries of the time-varying mixing matrix per TFM Results presented in Table 4 show that the sum of the absolute correlation values across the entries of the time-varying mixing matrix for each TFM, for associations of entries of the time-varying mixing matrix with the visual and motor task regressors. We observed that the sum of the correlation values was highest for TFM3 when related to the visual task regressor and for entries of TFM14 when related to the motor task regressor (for the selected dataset). These were the same TFMs for which the time series correlated most strongly to the task regressors.
| TFM | Sum of the absolute r-values | |
| Visual | Motor | |
| TFM1 | 1.33 | 0.92 |
| TFM2 | 1.20 | 1.03 |
| TFM3 | 3.83 | 1.69 |
| TFM4 | 1.16 | 1.10 |
| TFM5 | 1.74 | 1.35 |
| TFM6 | 1.59 | 2.04 |
| TFM7 | 1.09 | 0.98 |
| TFM8 | 1.53 | 1.80 |
| TFM9 | 0.96 | 0.92 |
| TFM10 | 0.80 | 0.86 |
| TFM11 | 0.54 | 1.03 |
| TFM12 | 1.22 | 1.49 |
| TFM13 | 1.35 | 0.95 |
| TFM14 | 1.38 | 2.33 |
| TFM15 | 0.94 | 0.93 |
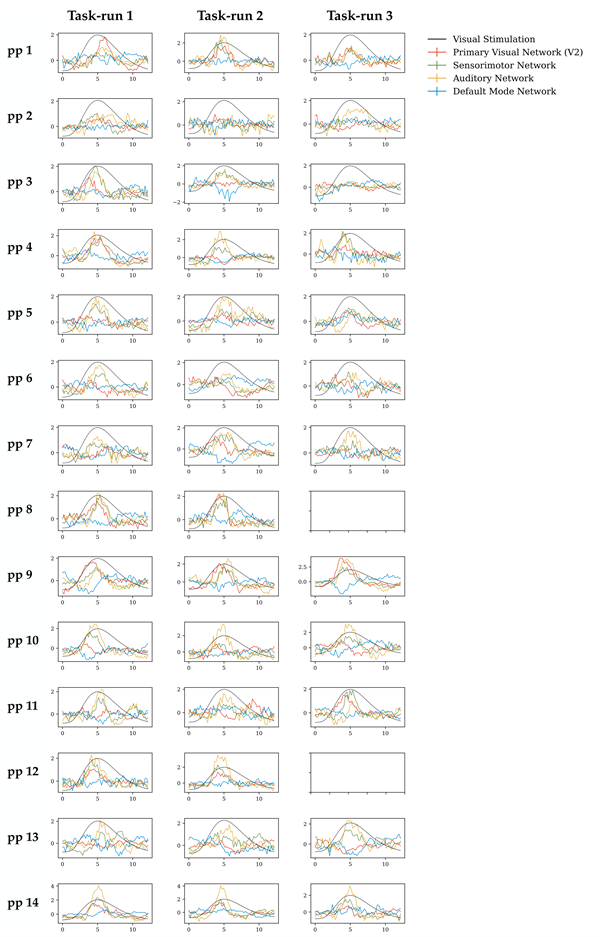
7. Stage 3 TV-TFM: Trial-averaged entries of the time-varying mixing matrix per task run, per-participant. We observed a pattern in the time-varying mixing matrix entries of the task-positive mode for the selected dataset that closely followed the task design. A similar trend was observed for most participants and task runs, which are presented in Figure 13. This figure shows the trial-averaged task-relevant entries of the time-varying mixing matrix for the TFM that was most strongly associated with the visual task regressor per task run, per participant.
Acknowledgements [back to top]
I would like to thank several people that made this thesis possible. First of all, I would like to thank Alberto, who not only came up with the initial idea and the derivation for this "Moonshine" method but with whom I went through all the ups and downs of the development process. Thank you for trusting me with it, for the endlessly interesting and fun discussions, and for deliberating the most complicated material with me at one moment and just as patiently explaining something basic the next. A major thanks also to Christian, who welcomed me into his lab group with open arms, who only needs a split second to hit the nail on the head, and gives spot-on feedback. Thank you for your concern for my wellbeing, facilitating Toolkits and courses, and for the fun talks. Thirdly, Nils: thank you for your consistency in meeting with me during quarantine, your feedback, and the interesting (project-related and other) discussions. I’m looking forward to the next four years. I would also like to thank my RetteRetraiteteam, Tim, Lotje, Christl, Gerwin and Yentl, who prevented lonely working days in quarantine and gave structure to my working days, and to Fonti who provided me with a working space and "oat cappuccinos" . Thank you Christl and Yentl for always being there, monitoring my well-being, and celebrating all highs and lows. I’d also like to kindly thank my parents, who have always supported me in many more ways than I can express in words, and who have taught me to use my abilities for a greater good. And last, but not least: Djarno, my love. Thank you for the wonderful, stable force you are in my life. For the desk you build me, and the endless meals you prepared. For smiling when my hyperfocus prevents me from paying attention to my immediate surroundings, or when I stumble over my own feet because my mind is solving problems. I couldn’t have done it, but more importantly, I wouldn’t have wanted to do it without all of you!
References [back to top]
Avants, B., Epstein, C., Grossman, M., & Gee, J. (2008). Symmetric diffeomorphic image registration with cross-correlation: Evaluating automated labeling of elderly and neurodegenerative brain. Medical Image Analysis, 12(1), 26–41.
Bandettini, P. A. (2009). What’s new in neuroimaging methods? Annals of the New York Academy of Sciences, 1156(1), 260–293.
Beckmann, C., Mackay, C., Filippini, N., & Smith, S. (2009). Group comparison of resting-state fMRI data using multi-subject ICA and dual regression. NeuroImage, 47, S148.
Beckmann, C., & Smith, S. (2004). Probabilistic independent component analysis for functional magnetic resonance imaging. IEEE Transactions on Medical Imaging, 23(2), 137–152.
Beckmann, C. F. (2012). Modelling with independent components. NeuroImage, 62(2), 891–901.
Beckmann, C. F., DeLuca, M., Devlin, J. T., & Smith, S. M. (2005). Investigations into
resting-state connectivity using independent component analysis. Philosophical Transactions of the Royal Society B: Biological Sciences, 360(1457), 1001–1013.
Bijsterbosch, J. (2017). Introduction to resting state fMRI functional connectivity. Oxford Neuroimaging Primers. Oxford, United Kingdom; New York, NY: Oxford University Press.
Bijsterbosch, J. D., Woolrich, M. W., Glasser, M. F., Robinson, E. C., Beckmann, C. F.,
Van Essen, D. C., Harrison, S.J. & Smith, S. M. (2018). The relationship between spatial configuration and functional connectivity of brain regions. eLife, 7:e32992.
Biswal, B., Zerrin Yetkin, F., Haughton, V. M., & Hyde, J. S. (1995). Functional connectivity in the motor cortex of resting human brain using echo-planar MRI. Magnetic Resonance in Medicine, 34(4), 537–541.
Cabral, J., Vidaurre, D., Marques, P., Magalhães, R., Silva Moreira, P., Miguel Soares, J., Deco, G., & Kringelbach, M. L. (2017). Cognitive performance in healthy older adults relates to spontaneous switching between states of functional connectivity during rest. Scientific Reports, 7(1), 1–13.
Calhoun, V., Adali, T., Pearlson, G., & Pekar, J. (2001). Spatial and temporal independent component analysis of functional MRI data containing a pair of task-related waveforms. Human Brain Mapping, 13(1), 43–53.
Cauley, S. F., Polimeni, J. R., Bhat, H., Wald, L. L., & Setsompop, K. (2014). Interslice leakage artifact reduction technique for simultaneous multislice acquisitions: Interslice leakage artifact reduction technique. Magnetic Resonance in Medicine, 72(1), 93–102.
Chang, C., & Glover, G. H. (2010). Time–frequency dynamics of resting-state brain connectivity measured with fMRI. NeuroImage, 50(1), 81–98.
Damoiseaux, J. S., Prater, K. E., Miller, B. L., & Greicius, M. D. (2012). Functional connectivity tracks clinical deterioration in Alzheimer’s disease. Neurobiology of Aging, 33(4), 828.e19–828.e30.
Damoiseaux, J. S., Rombouts, S. A. R. B., Barkhof, F., Scheltens, P., Stam, C. J., Smith, S. M., & Beckmann, C. F. (2006). Consistent resting-state networks across healthy subjects. Proceedings of the National Academy of Sciences, 103(37), 13848–13853.
Deco, G., Cabral, J., Woolrich, M. W., Stevner, A. B., van Hartevelt, T. J., & Kringelbach, M. L. (2017). Single or multiple frequency generators in on-going brain activity: A mechanistic whole-brain model of empirical MEG data. NeuroImage, 152, 538–550.
Deco, G., & Kringelbach, M. L. (2016). Metastability and coherence: Extending the communication through coherence hypothesis using a whole-brain computational perspective. Trends in Neurosciences, 39(3), 125–135.
Esteban, O., Markiewicz, C. J., Blair, R. W., Moodie, C. A., Isik, A. I., Erramuzpe, A., & Gorgolewski, K. J. (2019). fMRIPrep: A robust preprocessing pipeline for functional MRI. Nature Methods, 16(1):111–116.
Fonov, V., Evans, A., McKinstry, R., Almli, C., & Collins, D. (2009). Unbiased nonlinear average age-appropriate brain templates from birth to adulthood. NeuroImage, 47, S102.
Friston, K. J. (1998). Modes or models: A critique on independent component analysis for fMRI. Trends in Cognitive Sciences, 2(10), 373–375.
Friston, K. J., Frith, C. D., Liddle, P. F., & Frackowiak, R. S. J. (1993). Functional connectivity: The principal-component analysis of large (PET) data sets. Journal of Cerebral Blood Flow & Metabolism, 13(1), 5–14.
Glasser, M. F., Coalson, T. S., Bijsterbosch, J. D., Harrison, S. J., Harms, M. P., Anticevic, A., Van Essen, D. C., & Smith, S. M. (2018). Using temporal ICA to selectively remove global noise while preserving global signal in functional MRI data. NeuroImage, 181, 692–717.
Glerean, E., Salmi, J., Lahnakoski, J. M., Jääskeläinen, I. P., & Sams, M. (2012). Functional magnetic resonance imaging phase synchronization as a measure of dynamic functional connectivity. Brain Connectivity, 2(2), 91–101.
Gomez, D. E., Llera, A., Marques, J. P. F., Beckmann, C. F., & Norris, D. G. (2020).
Single-subject, single-session, temporal modes of brain activity. NeuroImage, 218, 116783.
Gorgolewski, K. J., Auer, T., Calhoun, V. D., Craddock, R. C., Das, S., Duff, E. P., & Poldrack, R. A. (2016). The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Scientific Data, 3(1), 1–9.
Greicius, M. D., Flores, B. H., Menon, V., Glover, G. H., Solvason, H. B., Kenna, & Schatzberg, A. F. (2007). Resting-state functional connectivity in major depression: Abnormally increased contributions from subgenual cingulate cortex and thalamus. Biological Psychiatry, 62(5), 429–437.
Greicius, M. D., Srivastava, G., Reiss, A. L., & Menon, V. (2004). Default-mode network activity distinguishes Alzheimer’s disease from healthy aging: Evidence from functional MRI. Proceedings of the National Academy of Sciences, 101(13), 4637–4642.
Greve, D. N. & Fischl, B. (2009). Accurate and robust brain image alignment using boundary-based registration. NeuroImage, 48(1), 63–72.
Hindriks, R., Adhikari, M., Murayama, Y., Ganzetti, M., Mantini, D., Logothetis, N., & Deco, G. (2016). Can sliding-window correlations reveal dynamic functional connectivity in resting-state fMRI? NeuroImage, 127, 242–256.
Hoge, W. S., Setsompop, K., & Polimeni, J. R. (2018). Dual-polarity slice-GRAPPA for concurrent ghost correction and slice separation in simultaneous multi-slice EPI. Magnetic Resonance in Medicine, 80(4), 1364–1375.
Hutchison, R. M., Womelsdorf, T., Allen, E. A., Bandettini, P. A., Calhoun, V. D., Corbetta, M., & Chang, C. (2013). Dynamic functional connectivity: Promise, issues, and interpretations. NeuroImage, 80, 360–378.
Hyvarinen, A. (1999). Fast and robust fixed-point algorithms for independent component analysis. IEEE Transactions on Neural Networks, 10(3), 626–634.
Iraji, A., Faghiri, A., Lewis, N., Fu, Z., Rachakonda, S., and Calhoun, V. (2020). Tools of the trade: Estimating time-varying connectivity patterns from fMRI data. PsyArXiv.
Jenkinson, M., Bannister, P., Brady, M., & Smith, S. (2002). Improved optimization for the robust and accurate linear registration and motion correction of brain images. NeuroImage, 17(2), 825–841.
Jenkinson, M., Beckmann, C. F., Behrens, T. E., Woolrich, M. W., & Smith, S. M. (2012). FSL. NeuroImage, 62(2), 782–790.
Kiviniemi, V., Vire, T., Remes, J., Elseoud, A. A., Starck, T., Tervonen, O., & Nikkinen, J. (2011). A sliding time-window ICA reveals spatial variability of the default mode network in time. Brain Connectivity, 1(4), 339–347.
Laird, A. R., Fox, P. M., Eickhoff, S. B., Turner, J. A., Ray, K. L., McKay, D. R., & Fox, P. T. (2011). Behavioral interpretations of intrinsic connectivity networks. Journal of Cognitive Neuroscience, 23(12), 4022–4037.
Lanczos, C. (1964). Evaluation of noisy data. Journal of the Society for Industrial and Applied Mathematics Series B Numerical Analysis, 1(1), 76–85.
Lehmann, M., Madison, C., Ghosh, P. M., Miller, Z. A., Greicius, M. D., Kramer, J. H., & Rabinovici, G. D. (2015). Loss of functional connectivity is greater outside the default mode network in nonfamilial early-onset Alzheimer’s disease variants. Neurobiology of Aging, 36(10), 2678–2686.
Leonardi, N. & Van De Ville, D. (2015). On spurious and real fluctuations of dynamic functional connectivity during rest. NeuroImage, 104, 430–436.
Liégeois, R., Laumann, T. O., Snyder, A. Z., Zhou, J., & Yeo, B. T. (2017). Interpreting temporal fluctuations in resting-state functional connectivity MRI. Neuroscience.
Liégeois, R., Li, J., Kong, R., Orban, C., Van De Ville, D., Ge, T., Sabuncu, M. R., & Yeo, B. T. T. (2019). Resting brain dynamics at different timescales capture distinct aspects of human behavior. Nature Communications, 10(1), 1–9.
Liu, X., & Duyn, J. H. (2013). Time-varying functional network information extracted from brief instances of spontaneous brain activity. Proceedings of the National Academy of Sciences, 110(11), 4392–4397.
Llera, A., Chauvin, R., Mulders, P., Naaijen, J., Mennes, M., & Beckmann, C. F. (2020). Spatial patterns for discriminative estimation. BioRxiv.
Lurie, D. J., Kessler, D., Bassett, D. S., Betzel, R. F., Breakspear, M., Kheilholz, S., & Calhoun, V. D. (2020). Questions and controversies in the study of time-varying functional connectivity in resting fMRI. Network Neuroscience, 4(1), 30–69.
Lustig, C., Snyder, A. Z., Bhakta, M., O’Brien, K. C., McAvoy, M., Raichle, M. E., & Buckner, R. L. (2003). Functional deactivations: Change with age and dementia of the Alzheimer type. Proceedings of the National Academy of Sciences, 100(24), 14504–14509.
Lynall, M. E., Bassett, D. S., Kerwin, R., McKenna, P. J., Kitzbichler, M., Muller, U., & Bullmore, E. (2010). Functional connectivity and brain networks in schizophrenia. Journal of Neuroscience, 30(28), 9477–9487.
Marques, J. P., Kober, T., Krueger, G., van der Zwaag, W., van de Moortele, P. F., & Gruetter, R. (2010). MP2RAGE, a self bias-field corrected sequence for improved segmentation and T1-mapping at high field. NeuroImage, 49(2), 1271–1281.
McKeown, M. (2003). Independent component analysis of functional MRI: What is signal and what is noise? Current Opinion in Neurobiology, 13(5), 620–629.
McKeown, M. J. & Sejnowski, T. J. (1998). Independent component analysis of fMRI data: Examining the assumptions. Human Brain Mapping, 6(5-6), 368–372.
Mulders, P., Llera, A., Tendolkar, I., van Eijndhoven, P., & Beckmann, C. (2018).
Personality profiles are associated with functional brain networks related to cognition and emotion. Scientific Reports, 8(1), 1–8.
Mulders, P. C., van Eijndhoven, P. F., Schene, A. H., Beckmann, C. F., & Tendolkar, I. (2015). Resting-state functional connectivity in major depressive disorder: A review. Neuroscience & Biobehavioral Reviews, 56, 330–344.
Niazy, R. K., Xie, J., Miller, K., Beckmann, C. F., & Smith, S. M. (2011). Spectral characteristics of resting state networks. Progress in Brain Research, 193, 259–276.
Nickerson, L. D., Smith, S. M., Öngür, D., and Beckmann, C. F. (2017). Using dual regression to investigate network shape and amplitude in functional connectivity analyses. Frontiers in Neuroscience, 11, 115.
Noteboom, S. (2018). Ultrafast fMRI for Studying Fine Hemodynamic Response Latency Differences (Master thesis, University of Twente and Donders Institute, the Netherlands).
Petridou, N., Gaudes, C. C., Dryden, I. L., Francis, S. T., & Gowland, P. A. (2013). Periods of rest in fMRI contain individual spontaneous events which are related to slowly fluctuating spontaneous activity. Human Brain Mapping, 34(6), 1319–1329.
Poldrack, R. A., Nichols, T., & Mumford, J. (2011). Handbook of Functional MRI Data Analysis. Cambridge, United Kingdom: Cambridge University Press.
Preti, M. G., Bolton, T. A., & Van De Ville, D. (2017). The dynamic functional connectome: State-of-the-art and perspectives. NeuroImage, 160, 41–54.
Ray, K. L., McKay, D. R., Fox, P. M., Riedel, M. C., Uecker, A. M., Beckmann, & Laird, A. R. (2013). ICA model order selection of task co-activation networks. Frontiers in Neuroscience, 7, 237.
Reuter, M., Rosas, H. D., & Fischl, B. (2010). Highly accurate inverse consistent registration: A robust approach. NeuroImage, 53(4), 1181–1196.
Rogers, B. P., Katwal, S. B., Morgan, V. L., Asplund, C. L., & Gore, J. C. (2010). Functional MRI and multivariate autoregressive models. Magnetic Resonance Imaging, 28(8), 1058–1065.
Rosa, M. J., Daunizeau, J., and Friston, K. J. (2010). EEG-fMRI integration: A critical review of biophysical modeling and data analysis approaches. Journal of Integrative Neuroscience, 9(4), 453–476.
Rosazza, C. and Minati, L. (2011). Resting-state brain networks: Literature review and clinical applications. Neurological Sciences, 32(5), 773–785.
Setsompop, K., Gagoski, B. A., Polimeni, J. R., Witzel, T., Wedeen, V. J., & Wald, L. L. (2012). Blipped-controlled aliasing in parallel imaging for simultaneous multislice echo planar imaging with reduced g-factor penalty. Magnetic Resonance in Medicine, 67(5), 1210–1224.
Singh, K. & Fawcett, I. (2008). Transient and linearly graded deactivation of the human default-mode network by a visual detection task. NeuroImage, 41(1), 100–112.
Smith, S. M., Fox, P. T., Miller, K. L., Glahn, D. C., Fox, P. M., Mackay, C. E., & Beckmann, C. F. (2009). Correspondence of the brain’s functional architecture during activation and rest. Proceedings of the National Academy of Sciences, 106(31), 13040–13045.
Smith, S. M., Jenkinson, M., Woolrich, M. W., Beckmann, C. F., Behrens, T. E., Johansen-Berg, H., & Matthews, P. M. (2004). Advances in functional and structural MR image analysis and implementation as FSL. NeuroImage, 23, 208–219.
Smith, S. M., Miller, K. L., Moeller, S., Xu, J., Auerbach, E. J., Woolrich, M. W., & Ugurbil, K. (2012). Temporally-independent functional modes of spontaneous brain activity. Proceedings of the National Academy of Sciences, 109(8), 3131–3136.
Tagliazucchi, E., Siniatchkin, M., Laufs, H., & Chialvo, D. R. (2016). The voxel-wise functional connectome can be efficiently derived from co-activations in a sparse spatio-temporal point-process. Frontiers in Neuroscience, 10, 381.
Treiber, J. M., White, N. S., Steed, T. C., Bartsch, H., Holland, D., Farid, N., & Chen, C. C. (2016). Characterization and correction of geometric distortions in 814 diffusion weighted images. PLoS one, 11(3), e0152472.
Tustison, N. J., Avants, B. B., Cook, P. A., Zheng, J., Egan, A., Yushkevich, P. A., & Gee,
J. C. (2010). N4ITK: Improved N3 bias correction. IEEE Transactions on Medical Imaging, 29(6), 1310–1320.
Uddin, L. Q., Supekar, K., & Menon, V. (2010). Typical and atypical development of functional human brain networks: Insights from resting-state fMRI. Frontiers in Systems Neuroscience, 4.
van Oort, E., Koopmans, P., Boyacioglu, R., Barth, M., Beckmann, C., & Norris, D. (2012). Frequency characteristics of large scale resting state networks using 7T Spin Echo EPI. Proceedings of the International Society for Magnetic Resonance in Medicine, 21.
van Oort, E. S., Mennes, M., Schröder, T. N., Kumar, V. J., Jimenez, N. I. Z., Grodd, W., Doeller, C.F. & Beckmann, C. F. (2018). Functional parcellation using time courses of instantaneous connectivity. NeuroImage, 170, 31–40.
Vidaurre, D., Quinn, A. J., Baker, A. P., Dupret, D., Tejero-Cantero, A., & Woolrich, M. W. (2016). Spectrally resolved fast transient brain states in electrophysiological data. NeuroImage, 126, 81–95.
Vidaurre, D., Smith, S. M., & Woolrich, M. W. (2017). Brain network dynamics are hierarchically organized in time. Proceedings of the National Academy of Sciences, 114(48), 12827–12832.
Wang, S., Peterson, D. J., Gatenby, J. C., Li, W., Grabowski, T. J., & Madhyastha, T. M. (2017). Evaluation of field map and nonlinear registration methods for correction of susceptibility artifacts in diffusion MRI. Frontiers in Neuroinformatics, 11.
Yaesoubi, M., Allen, E. A., Miller, R. L., & Calhoun, V. D. (2015). Dynamic coherence analysis of resting fMRI data to jointly capture state-based phase, frequency, and time-domain information. NeuroImage, 120, 133–142.
Zhang, Y., Brady, M., & Smith, S. (2001). Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Transactions on Medical Imaging, 20(1), 45–57.
